John L. McCracken

Research
Biological Electron Paramagnetic Resonance
(Research Description PDF)
Electron Paramagnetic Resonance (EPR) spectroscopy provides an ideal tool for the determination of the structures of paramagnetic centers in chemical systems. The origin of this structural information is the spin-spin coupling between the magnetic moments of the paramagnetic center and nuclei that lie less than 6 Å away. Unfortunately, these spin-spin couplings are often weak and as such, they are buried by the inhomogeneous broadening of the EPR absorbtion lineshape. In the McCracken lab, we are applying the advanced EPR methods of Electron Spin Echo Envelope Modulation (ESEEM) and Electron-Nuclear Double Resonance (ENDOR) to determine the structures about paramagnetic centers in metalloenzymes. Our studies are aimed at using the information we gain from these experiments to understand the chemistry that occurs at metal centers and answer questions concerning structure-function relationships that cannot be addressed using other structural tools like NMR or X-ray crystallography in isolation.
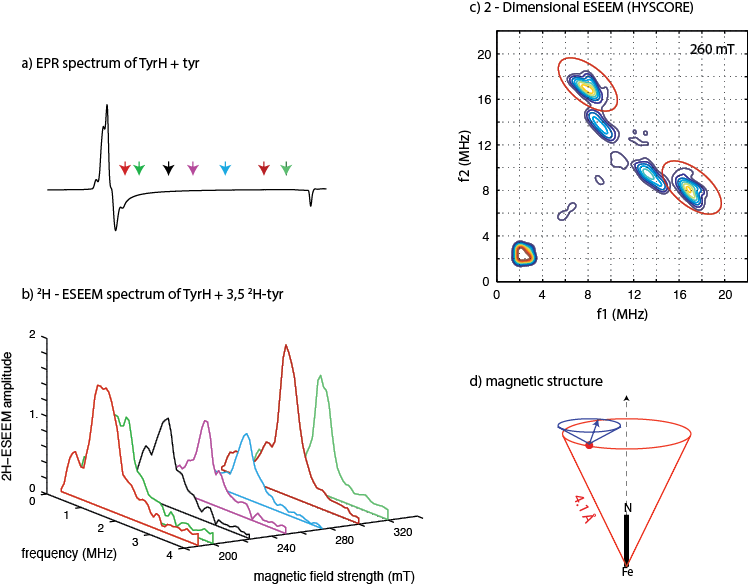
The figure shown below details two different applications of ESEEM spectroscopy to characterize the ligation structure of an Fe(II) ion located at the heart of the catalytic site of the enzyme Tyrosine Hydroxylase. This enzyme is present in the central nervous system of mammals and catalyzes the rate-limiting step in the biosynthesis of the catecholamine neurotransmitters, dopamine, epinephrine and norepinephrine. Our gateway into the structure is the EPR spectrum of an {FeNO}7 derivative of the enzyme and is shown in figure (a). This spectrum is about 200 mT wide and provides no features that can be attributed to ligands bound to Fe(II), or substrates and cofactors that may bind to the enzyme in Fe(II)’s second coordination sphere. Figure (b) shows 2H-ESEEM spectra, obtained at seven different magnetic field positions across the EPR spectrum, that arise from hyperfi coupling between a deuterium atom of 3,5 2H tyrosine bound to the enzyme and the paramagnetic Fe-NO center. The amplitudes and lineshapes of these spectra can be fit to a spin Hamiltonian model to provide the location of the coupled deuteron with respect to the axis of the Fe-NO bond, and the direction of the C-2H bond associated with the labeled substrate. Figure (d) summarizes these results showing that substrate tyrosine binds so that a coupled deuteron (red ball in figure d) is positioned Å from the Fe(II) and that the vector connecting the metal ion with this coupled deuteron makes an angle of 25° with the Fe-NO bond axis. These data represent the fi st structural information gained on the binding of the amino acid substrate at the catalytic site of this family of enzymes. By repeating these measurements on substrates deuterated at other positions, our crude magnetic structure can be built into an atomic level structure. The second type of experiment is a 2-dimensional ESEEM measurement that has proved useful for viewing the stronger hyperfine couplings that arise from the Fe(II) ligands. The spectrum shown in figure (c) was collected at 260 mT (aqua arrow in figure(a)) and shows off-diagonal cross-peaks, circled in red, that are diagnostic for bound water and/ or hydroxide ligands.
Area(s) of Interest
Physical (Ph)
Biological (Bi)
Selected Publications
Characterization of Water Coordination to Ferrous Nitrosyl Complexes with fac– N2O, cis–N2O2 and N2O3 Donor Ligands, McCracken, J.; Cappillino, P.J.; McNally, J.S.; Krzyaniak, M.D.; Howart, M.; Tarves, P.C.; Caradonna, J.P.; Inorganic Chemistry, 2015, 54, 6486–6497.
Structural Characterization of the Catalytic Sites of Mononuclear Non-Heme Iron Hydroxylases Using 2H-ESEEM, McCracken, J.; in Peter Qin, Kurt Warncke, eds.: Electron Paramagnetic Resonance Investigations of Biological Systems by Using Spin Labels, Spin Probes, and Intrinsic Metal Ions, Part A, Vol 563, MIE, UK: Academic Press, 2015, pp. 285-309.
Spectroscopic analysis of 2-oxoglutarate-dependent oxygenases: TauD a case study, Proshlyakov, D.A.; McCracken, J; Hausinger, R.P.; J. Biol. Inorg. Chem. 2017, 22, 367 – 379.
The Lactate Racemase Nickel-Pincer Cofactor Operates by a Proton-Coupled Hydride Transfer Mechanism, Rankin, J.A.; Mauban, R.C.; Fellner, M.; Desguin, B.; McCracken, J.; Hu, J.; Varganov, S.A.; Hausinger, R.P.; Biochemistry 2018, 57, 3244-3251.
Quasi-Monodisperse Transition Metal-Doped BaTiO3 (M=Cr, Mn, Fe, Co) Colloidal Nanocrystals with Multiferroic Properties, Costanzo, T.; McCracken, J.; Caruntu, G.; Rotaru, A.; ACS Applied Nano Materials 2018, 1, 4863-4874.
Synthesis and Characterization of a neutral U(II) Arene Sandwich Complex, Billow, B.; Livesay, B.N.; Mokhtarzadeh, C.G., McCracken, J.; Shores, M.P.; Boncella, J.M.; Odom, A.L.; J. Am. Chem. Soc. 2018, 140, 17369 – 17373.
Electronic and Structural Comparisons between Iron(II/III) and Ruthenium(II/III) Imide Analogues, Aldrich, K.E.; Fales, B.S.; Singh, A.K.; Staples, R.J.; Levine, B.G.; McCracken, J.; Smith, M.R.; Odom, A.L.; Inorganic Chemistry 2019, 58, 11699.
CV
B.S., 1978, Univ. of Illinois
Ph.D., 1983, Univ. of California, Berkeley
Postdoctoral Fellow, 1983-85, Albert Einstein College of Medicine
Associate Director, 1985-89, NIH Pulsed EPR Research Resource
Curriculum Vitae (PDF)
Awards
| Year | Award | Organization |
|---|---|---|
| 2012 | CNS Distinguished Faculty Award | |
| 2012 | Outstanding Faculty Award | College of Natural Science |
| 2000 - 2002 | Member | National Institutes of Health (NIH Special Study Sections) |
| 1997 | University Teacher-Scholar Award | Michigan State University |
| 1996 - 2000 | Member | National Institutes of Health (NIH Metallobiochemistry Study Section) |
| 1995 - 1996 | Member | National Institutes of Health (NIH Special Study Sections) |
| 1995 | Teacher-Scholar Award | Michigan State University (College of Natural Science) |
| 1992 - 2004 | Officer, MSU Local Section | American Chemical Society |
| 1990 - 1991 | Member | National Institutes of Health (NIH Special Study Sections) |
| 1985 - 1999 | NIH National Pulsed EPR Research Resource Governing Board | National Institutes of Health |
| 1983 | Ph.D. | University of California, Berkeley |
| 1981 | Chancellor's Fellow in Chemistry | University of California, Berkeley |
| 1978 | Bachelor of Science with Highest Distinction in Chemistry | University of Illinois, Urbana-Champaign |
| 1974 - 1978 | Edmund James Scholar | University of Illinois, Urbana-Champaign |
| Recognition of many contributions to magnetic resonance spectroscopy | Miami University of Ohio | |
| Chairperson | Michigan State University (Department of Chemistry) |